The Library in Atomes
The Atomes program includes a library with a number of molecules to be used when building or editing atomistic models:
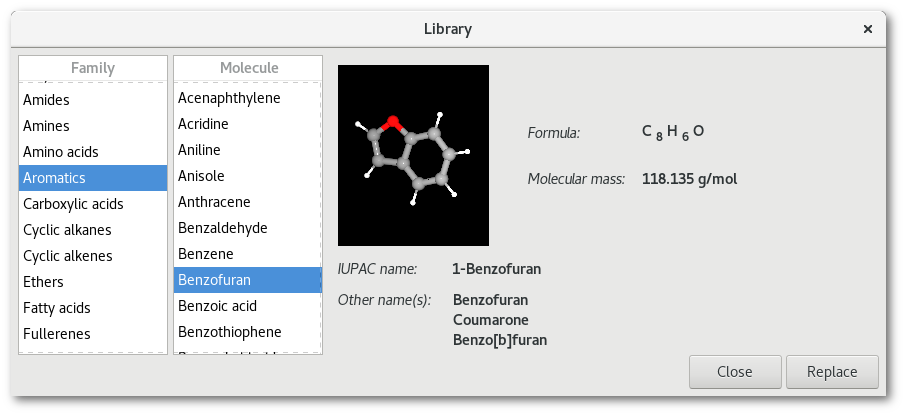
Available molecules can be found in the library by sorted in the following families:
Misc
Alcohols
Aldehydes
Alkanes
Alkenes
Alkynes
Amides
Amines
Amino acids
Aromatics
Carboxylic acids
Cyclic alkanes
Cyclic alkenes
Ethers
Fatty acids
Fullerenes
Heterocyclics
Macrocycles
Ketones
Nitriles
Nucleobases
Steroids
Sugars (Linears)
Sugars (Cyclics)
Sulfoxides
Thiols
Each time a new molecule is selected in the tree view on the left side of the window (see figure 3.1), then the right side of the "Library" dialog is refreshed with the data of the newly selected molecule. The data includes a 3D representation included in an active OpenGL window that allows to rotate the molecule:
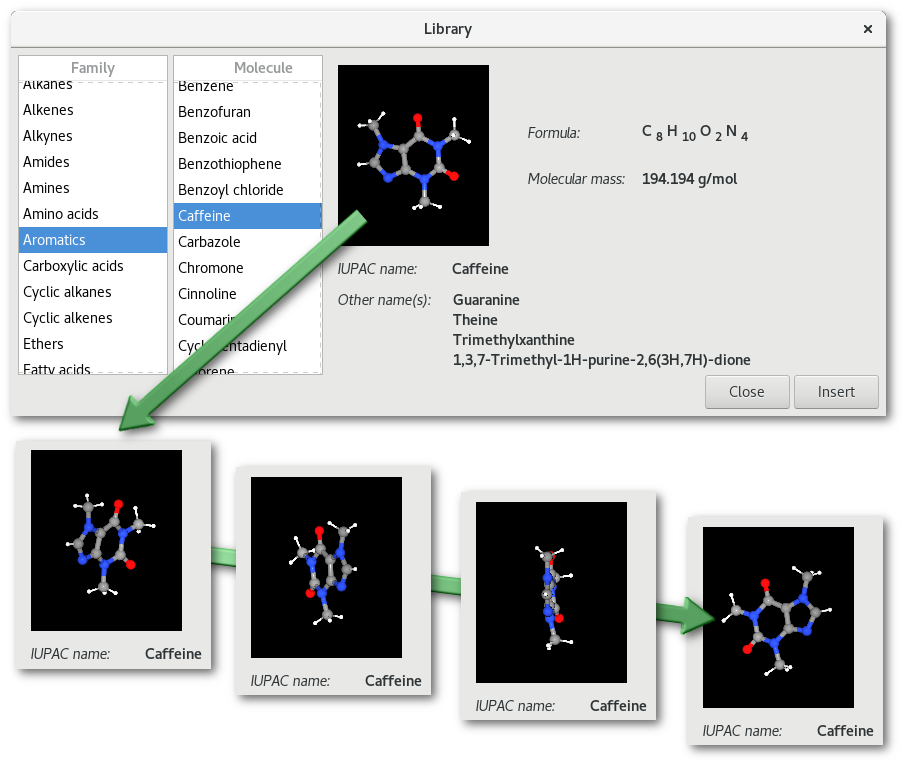
The data files that contain the information regarding each molecule in the library are located in the "bin/library/molecules" directory.
To each molecule that appears in the "Library" tree view, a file with the ".sml" extension can be found in the corresponding family subfolder.
For the example in figure 3.2:
The family is: "
Aromatics"The molecule is: "
Caffeine"The corresponding file is: "
Aromatics/caffeine.sml".
New files can be added to the library to be found by Atomes at startup, providing that the new files are added to one of the families of the library. These files must have the ".sml" extension, follow the "XML" encoding rules, and the following structure:
The "SCL" "Simple chemical library XML file" format
<?xml version="1.0" encoding="UTF-8"?>
<!– Simple chemical library XML file –>
<scl-xml>
<class>Aromatics</class>
<names>
<library-name>Caffeine</library-name>
<iupac-name>Caffeine</iupac-name>
<other-names>
<name>Guaranine</name>
<name>Theine</name>
<name>Trimethylxanthine</name>
<name>1,3,7-Trimethyl-1H-purine-2,6(3H,7H)-dione</name>
</other-names>
</names>
<chemistry>
<atoms>24</atoms>
<species number="4">
<label id="0" num="8">C</label>
<label id="1" num="10">H</label>
<label id="2" num="2">O</label>
<label id="3" num="4">N</label>
</species>
</chemistry>
<coordinates>
<atom id="1" sp="0" x="1.785021" y="-0.779129" z="-0.255949"/>
<atom id="2" sp="0" x="0.401929" y="1.318216" z="-0.016004"/>
<atom id="3" sp="0" x="-0.733464" y="0.413478" z="0.038347"/>
<atom id="4" sp="0" x="-0.617909" y="-0.973998" z="-0.109571"/>
<atom id="5" sp="0" x="-2.772439" y="-0.562269" z="0.209813"/>
<atom id="6" sp="0" x="0.735608" y="-3.069196" z="-0.210904"/>
<atom id="7" sp="0" x="-2.730788" y="1.961282" z="0.459758"/>
<atom id="8" sp="0" x="2.908891" y="1.458603" z="-0.240195"/>
<atom id="9" sp="1" x="-3.848274" y="-0.720508" z="0.332764"/>
<atom id="10" sp="1" x="1.561100" y="-3.446348" z="-0.829847"/>
<atom id="11" sp="1" x="-0.192018" y="-3.562176" z="-0.530166"/>
<atom id="12" sp="1" x="0.935512" y="-3.340819" z="0.835372"/>
<atom id="13" sp="1" x="-2.333498" y="2.706064" z="-0.245137"/>
<atom id="14" sp="1" x="-2.525360" y="2.320312" z="1.478733"/>
<atom id="15" sp="1" x="-3.818507" y="1.892617" z="0.326144"/>
<atom id="16" sp="1" x="3.674681" y="0.991531" z="-0.874770"/>
<atom id="17" sp="1" x="3.303682" y="1.537341" z="0.782105"/>
<atom id="18" sp="1" x="2.721889" y="2.471199" z="-0.622647"/>
<atom id="19" sp="2" x="2.892437" y="-1.297551" z="-0.212698"/>
<atom id="20" sp="2" x="0.363733" y="2.532611" z="0.115710"/>
<atom id="21" sp="3" x="1.659632" y="0.660920" z="-0.271271"/>
<atom id="22" sp="3" x="0.612806" y="-1.608906" z="-0.393730"/>
<atom id="23" sp="3" x="-2.109760" y="0.657267" z="0.234906"/>
<atom id="24" sp="3" x="-1.874903" y="-1.560540" z="-0.000764"/>
</coordinates>
</scl-xml>