The Model menu
The "Atom(s)" menu
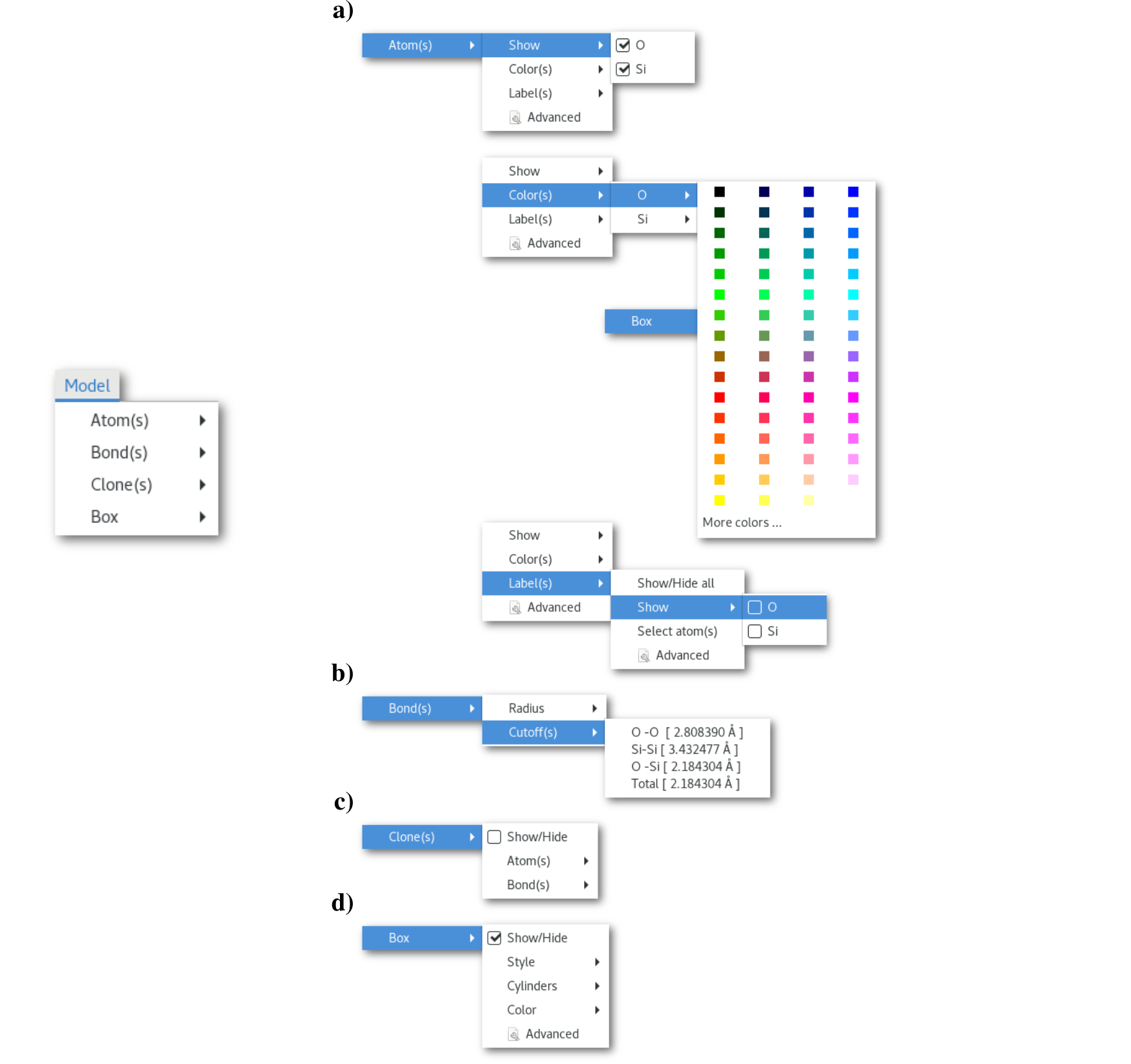
The "Atom(s)" menu [Fig. 5.5-a] offers to configure some visual features related to the chemical species in the model. This menu contains shortcuts for some the properties that can be adjusted using the more advanced "Atom(s) configuration" dialog, and that can be opened using any of the "Advanced" buttons in the menu. The "Atom(s) configuration" dialog is presented in figure 5.6:
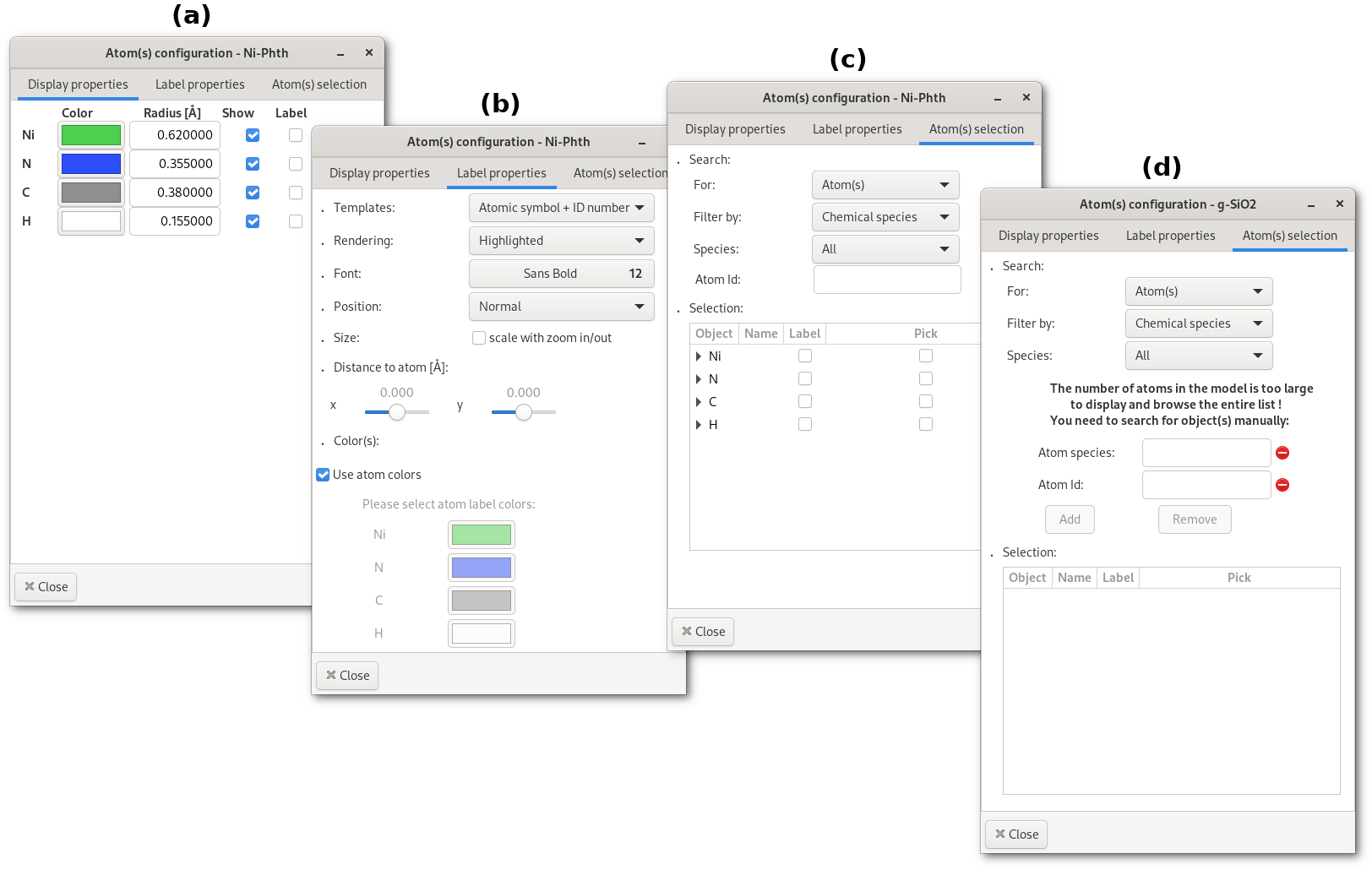
The window contains a notebook with 3 tabs:
(a) The "
Display properties" tab: to adjust chemical species colors, atomic labels, and depending on the model style:Ball and sticks: atomic radius
Wireframe or Points: point size
(b) The "
Label properties" tab: to adjust the aspect of the atomic labels.(c) or (d) The "
Atom(s) selection": to search for atom(s) in the model. If the model contains less than 10 000 atoms the entire list of atoms is displayed (c), otherwise it becomes too complicated to display and a search engine is proposed (d).
The "Bond(s)" menu
Via the "Bond(s)" menu [Fig. 5.5-b] it is possible to adjust the bond cutoff(s), and depending on the model style:
Ball and sticks: bonds radius(ii)
Cylinders: bond radius
Wireframe: line width
The "Clone(s)" menu
The Atomes program uses clones to illustrate the presence of atomic bond(s) on the edges of the simulation box and existing because of the periodic boundary conditions (see section 5.1 for more information on the PBC).
For the "Clone(s)" menu to be activated two conditions must be fulfilled: to use PBC, ie. the model must have a periodicity, and, for such bonds via PBC to exist. If both conditions are met, then the first button of the menu to "Show/Hide Clones" will be active and it becomes possible to show or hide the clones.
Please note that clone atoms/bonds are translucent to make them easily distinguishable from their atom/bond counterparts, figure 5.7 illustrates the clones idea using the silica glass model:
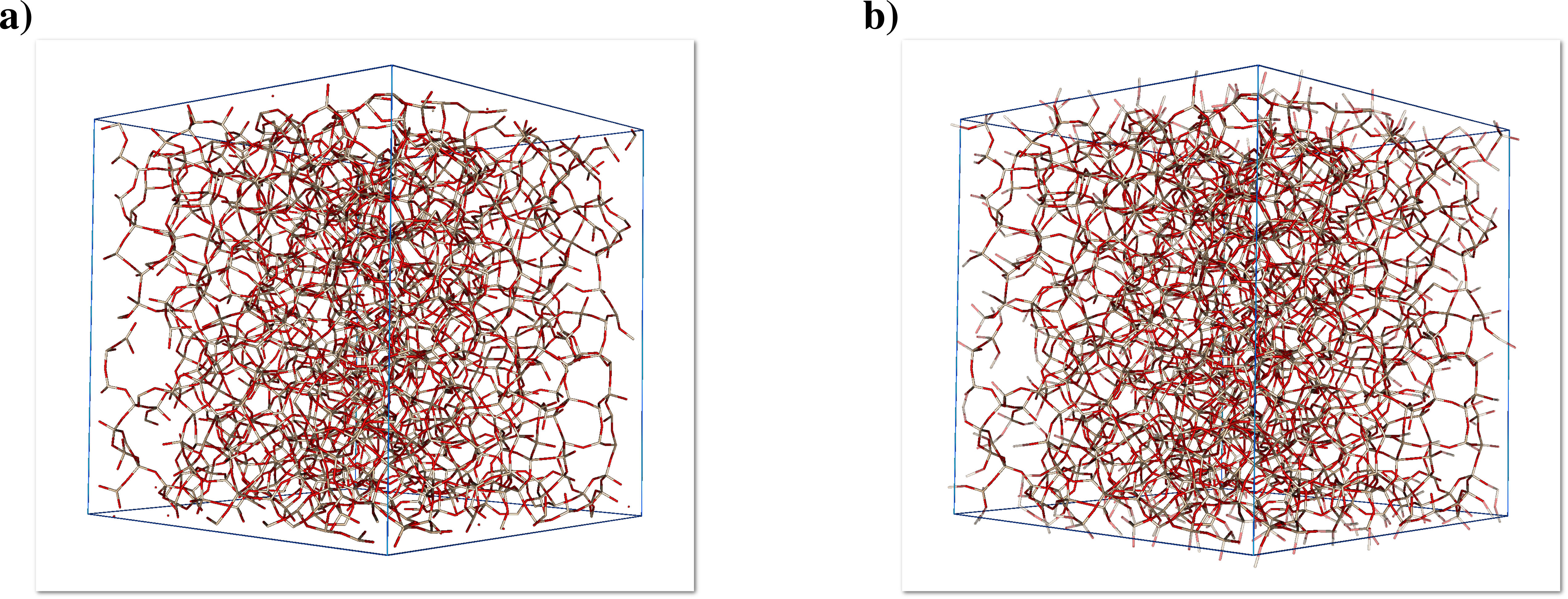
If the clones are visible then the two "Atom(s)" and "Bond(s)" submenus will be activated as well. These menus are reproductions of the upper level "Atom(s)" and "Bond(s)" menus, and windows, but dedicated to the clones, this allows to make the clones even more distinguishable in the 3D representation.